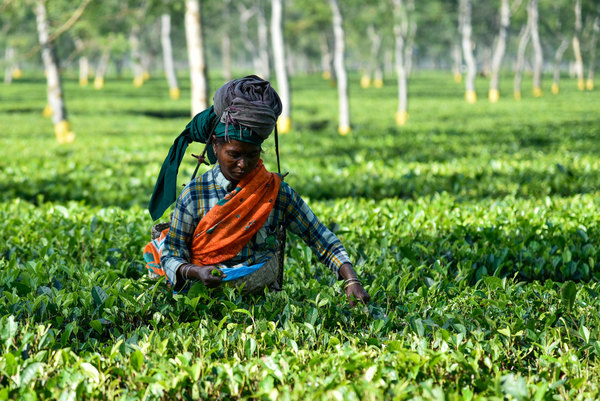
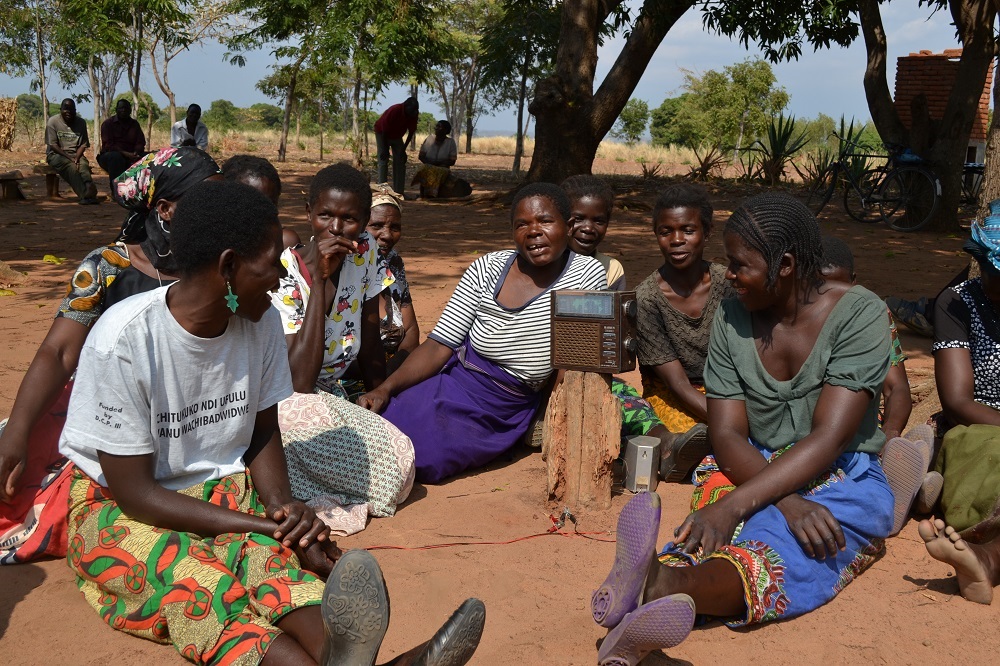How comics, pop music and drama deliver down-to-earth messages to help African farmers improve their soil
5 December is World Soil Day. Soil is a life source for healthy ecosystems, healthy food and, ultimately, healthy humans. With a growing population, the soil that’s cultivated will have to feed many more people, but it will only be able to do this if soil fertility is managed.
E-Learning Course on Bioinformatics of Animal Viruses
Nucleotide sequencing has become a very popular technique for diagnosis and characterization of pathogens and is accessible to most veterinary practices. A nucleotide sequence provides information on the nature of the pathogen, its source and its main characteristics such as strain, virulence and drug resistance. Bioinformatics provides tools to gather, store, and analyse these biological…
African Swine Fever on the Move – China, the EU and FAO Assessing Preparedness in East and Southeast Asia, the Region with >50% of the World Pig Population
By M Djuric, DVM African Swine Fever (ASF) continues to spread in traditionally endemic sub-Saharan Africa, but it is also expanding into previously ASF-free countries with a new front opening up along the Caucasus and Eastern Europe. The risk of ASF entering China is of particular concern since the country keeps almost half of the…
Global Milk Production Increasing at Fast Pace
By Miroslav Djuric, Editor (Dairy Science Abstracts) Global milk production in 2012 is forecast to reach 760 million tonnes, according to a new report published in the Food Outlook by the United Nation’s Food and Agriculture Organisation (FAO). This would represent an annual increase of 3%, largely due to the increased production in Asia, Oceania…
Recent developments in the world of biofuels
Opinions on the use of crops for biofuel and bioenergy continue to be polarized – are they a ‘good thing’ or not? When are they a ‘good thing’? Who benefits? How do you measure the impacts and their interactions at a local, national and international level on food security, land resources, water, greenhouse gas emissions,…
Biodiversity – The more the merrier!
May 22nd is the International Day for Biological Diversity 2008. This year’s theme is ‘Biodiversity in Agriculture‘. According to the Convention on Biodiversity who are co-promoting the day’s festivities along with such luminaries of food and nutrition as the FAO, modern food production is responsible for both increasing and decreasing biodiversity. One of the things…